IICD Researchers Introduce Starfysh, a Breakthrough Tool Transforming Spatial Gene Expression Analysis
IICD researcher Elham Azizi - Herbert and Florence Irving Assistant Professor of Cancer Data Research and Assistant Professor of Biomedical Engineering - and her team introduce an innovative computational tool named Starfysh, designed to revolutionize the study of gene expression within tissues. This breakthrough has far-reaching implications because cells in our bodies can behave differently depending on their context and surroundings, and characterizing these differences can help us understand tissue development, function, and disease response, particularly in contexts such as cancer.
The study was published in Nature Biotechnology. The three graduate students and co-first authors - Siyu He, Yinuo Jin, and Achille Nazaret - played a pivotal role in bringing this project to fruition. Starfysh is also the result of an interdisciplinary collaboration with IICD research groups of José McFaline-Figueroa and David Blei, researchers at Columbia Biomedical Engineering - Kam Leong’s group - and Memorial Sloan Kettering Cancer Center - George Plitas and Alexander Rudensky’s groups.
"Starfysh represents a paradigm shift in spatial gene expression analysis, providing researchers with unprecedented insights into tissue organization and cellular dynamics," says Jin, a PhD student in biomedical engineering (Azizi’s lab) and co-author of the study.
At the heart of Starfysh lies its ability to unravel the complexities of gene expression within different tissue environments. Traditional analytical tools often struggle to dissect refined cell types or compare spatial organization across multiple tissues. Azizi's team recognized these limitations and set out to develop a solution that could overcome them.

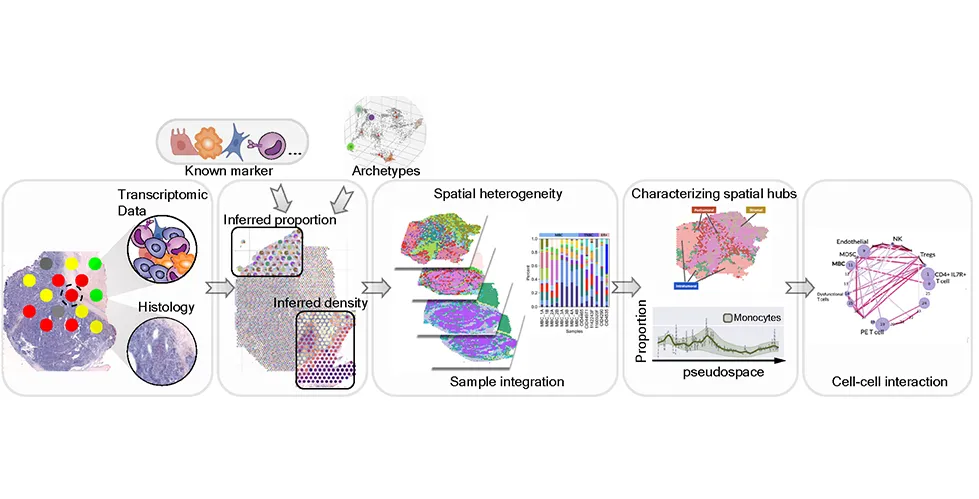
Starfysh combines spatially-resolved gene expression data with histology images, providing researchers with a comprehensive view of tissue structure and cellular activity. “By employing advanced machine learning techniques and mathematical concepts like archetypal analysis, the tool can identify various cell types and states without the need for single-cell resolution references,” explains He, a former PhD student in biomedical engineering (Azizi/Leong’s labs) and co-author of the study.
“One of the key challenges addressed by Starfysh is the spatial distribution of immune cells in aggressive subtypes of breast cancer,” adds Nazaret, a PhD student in computer science (Azizi/Blei’s labs) and co-author of the study. “By mapping out these patterns, researchers can gain insights into why certain immune cell types fail to infiltrate therapy-resistant tumors.” This understanding could pave the way for more effective treatment strategies tailored to combat resistant forms of cancer.
The Azizi lab's expertise in biomedical engineering and cancer data research has been instrumental in the development of Starfysh. Their work focuses on characterizing the complex populations of interacting cell types within and across tumor microenvironments. By leveraging cutting-edge single-cell genomic technologies and machine learning methods, Azizi aims to decipher the underlying cancer progression and immune dysfunction.
Azizi and her team are already looking ahead to the future possibilities enabled by Starfysh. “From constructing atlases of healthy and diseased tissues to guiding personalized cancer therapies, the potential applications of this computational tool are vast and promising,” explains Azizi.
The acceptance of their paper in Nature Biotechnology marks a significant milestone for Azizi and her team. Their groundbreaking research not only showcases the potential of Starfysh in advancing tissue analysis but also underscores Columbia University's commitment to pushing the boundaries of scientific discovery.